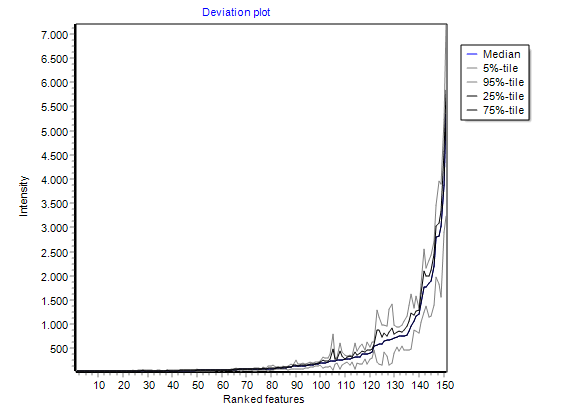
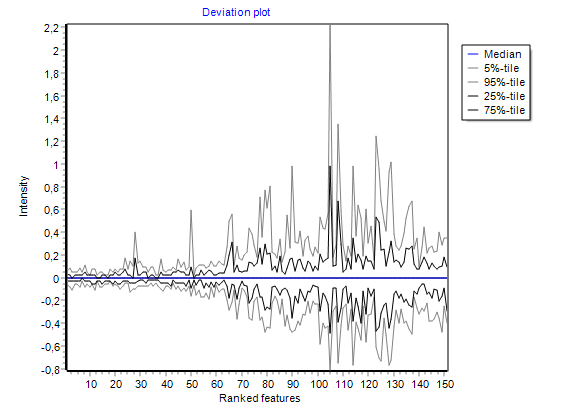
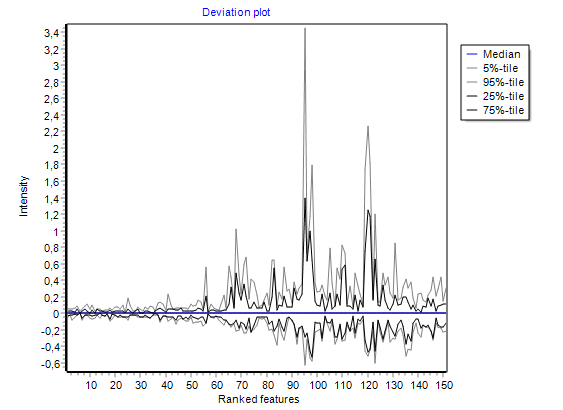
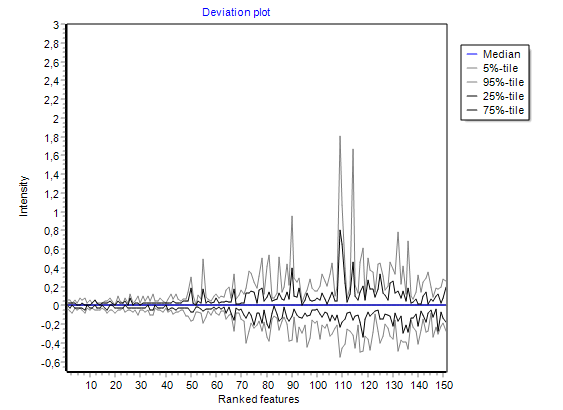
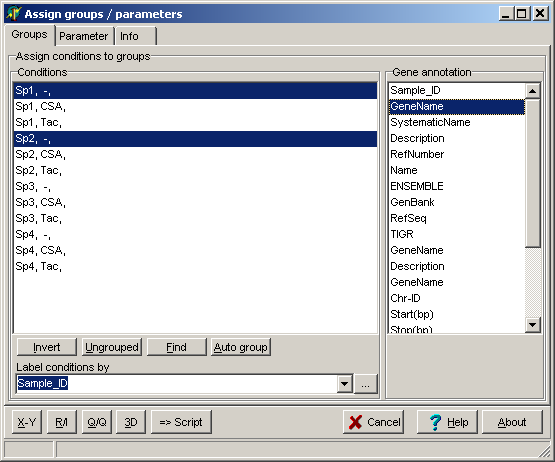
visualising intensities / ratios
of the expression matrix.
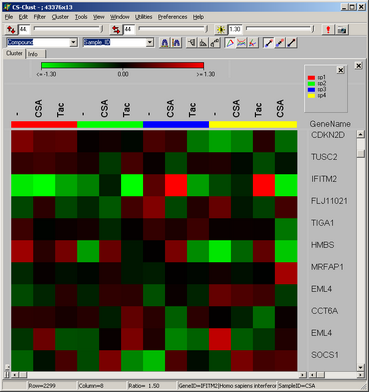
The famous red-green heat maps
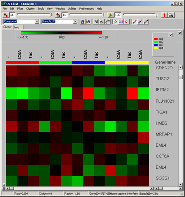
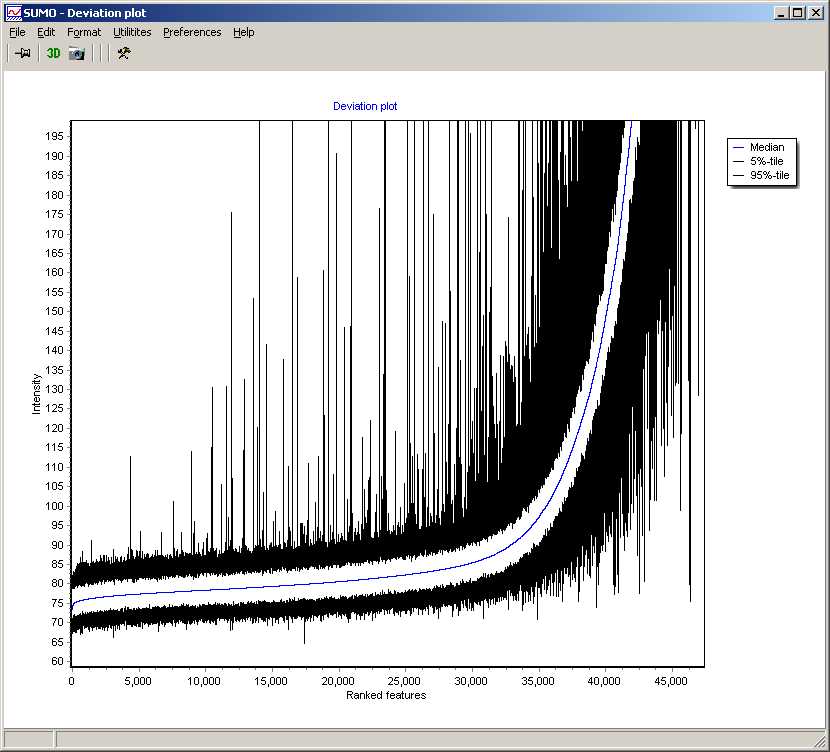
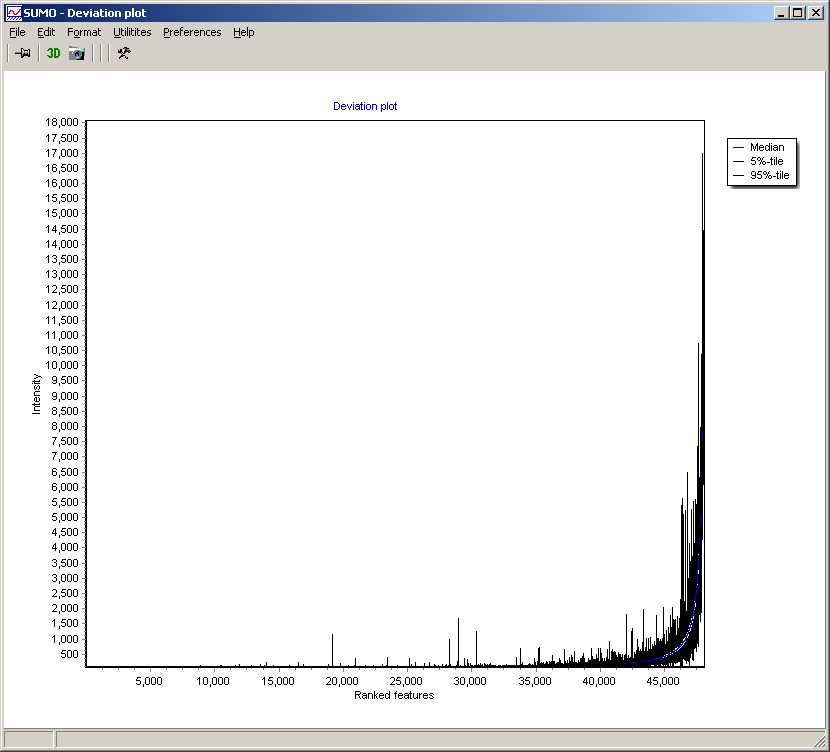
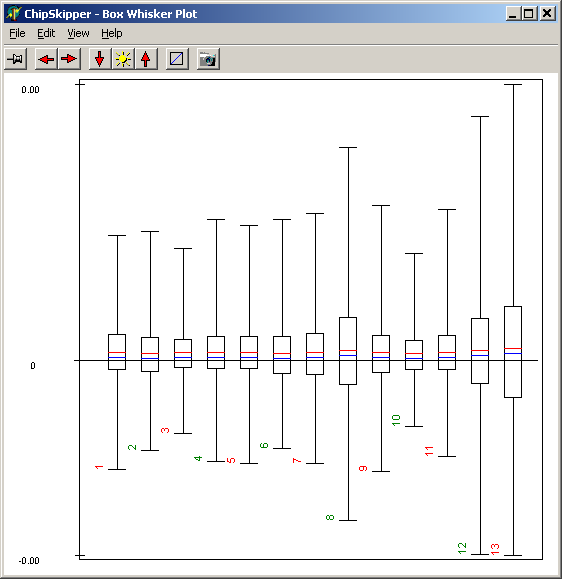
summarizing global intensity/ratio distribution
in all hybridisations
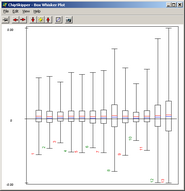
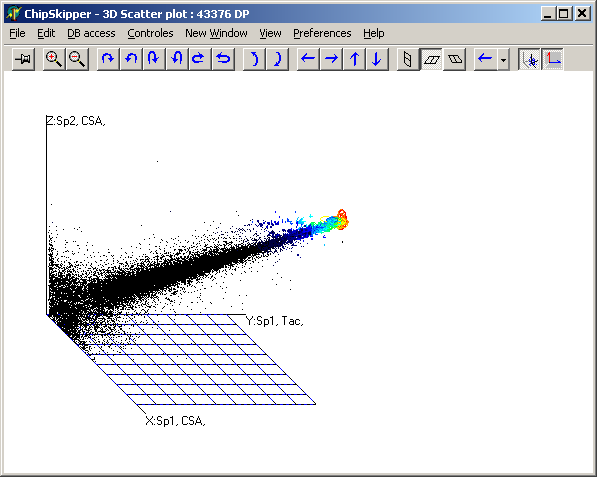
Analyze pair wise signal distributions
See more details about the scatterplot viewer.
of hybridisations
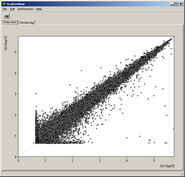
Get an overview of signal distributions
in all your sample data
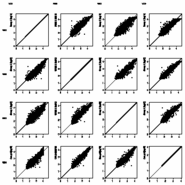
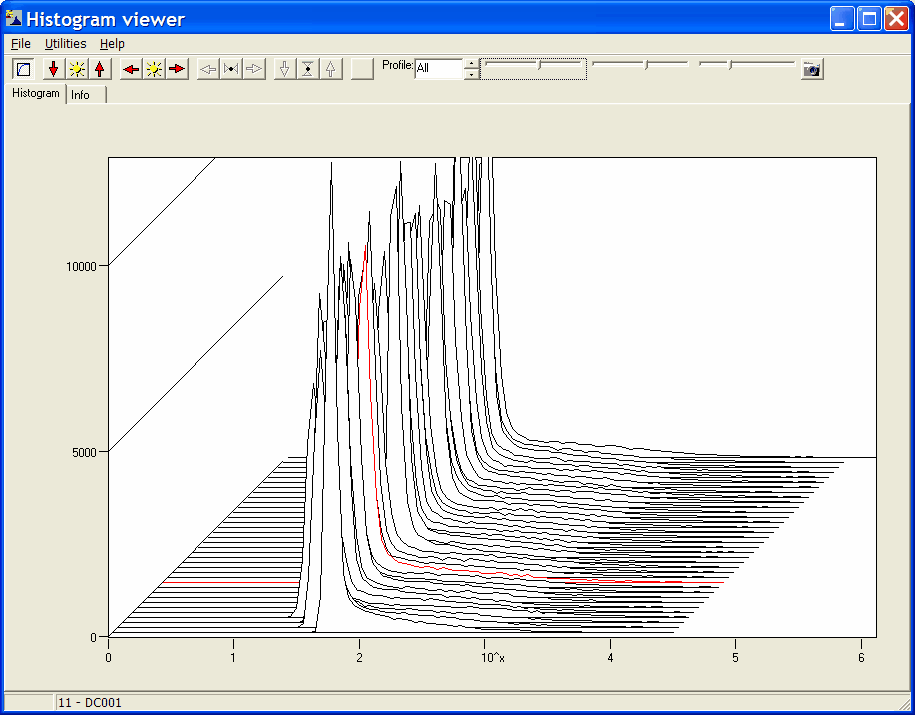
Show signal distribution in any
or all loaded samples
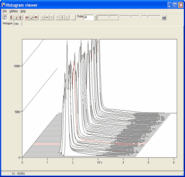
Illustrating e.g. gene/sample
data profiles
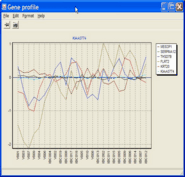
Illustrating inter-sample
similarity
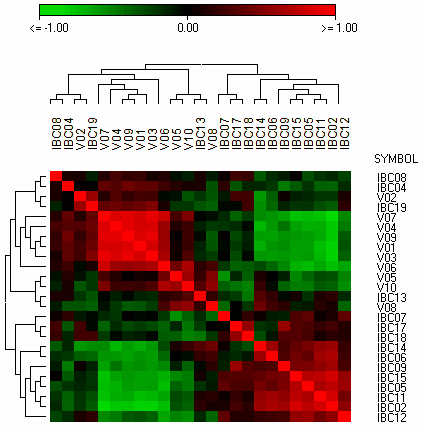
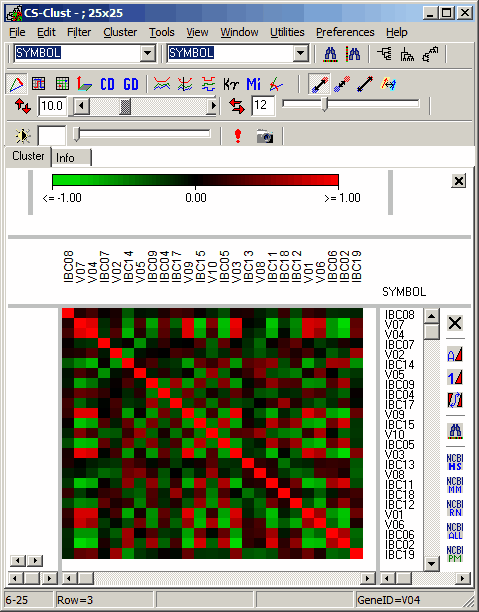
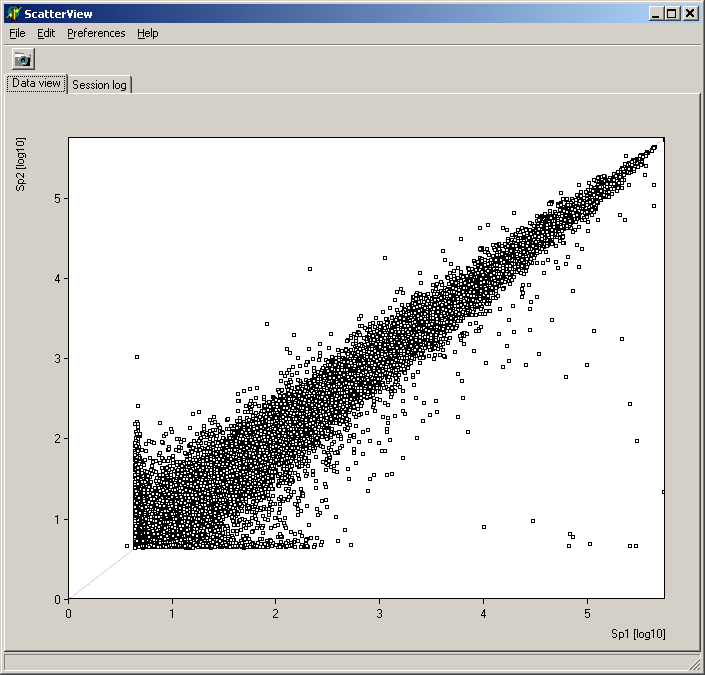
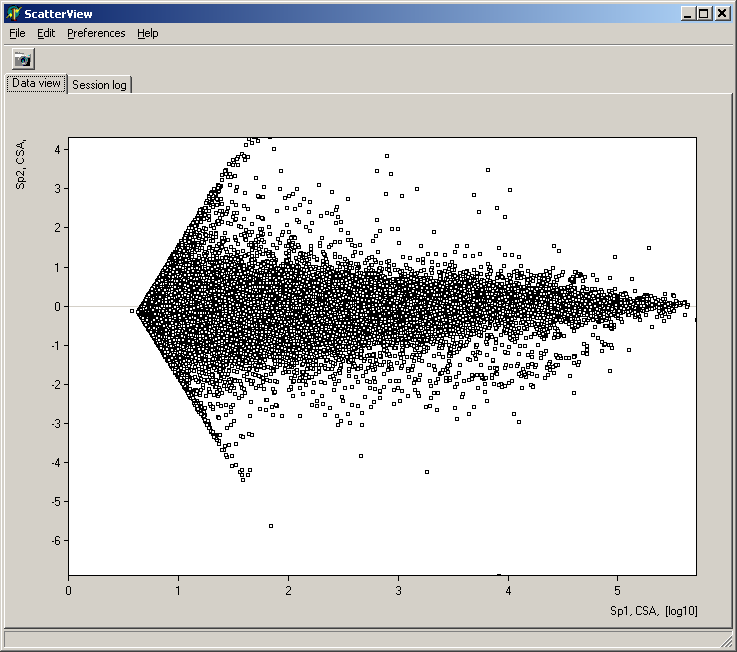
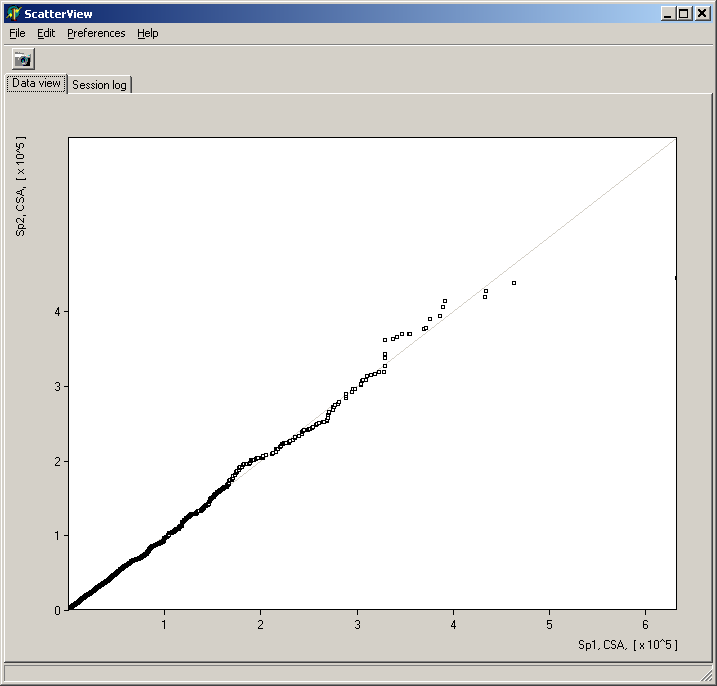 See more details about the Scatterplot viewer
See more details about the Scatterplot viewer