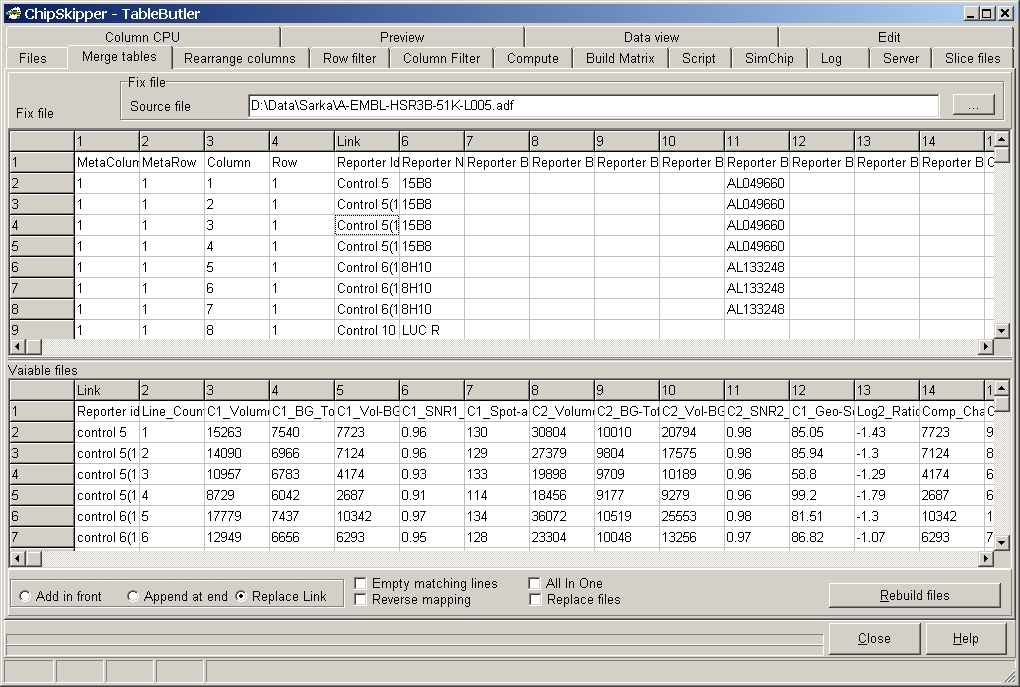
This function allows to merge tables via a Link column. Depending on the content of the datafield in the Link column, the corresponding line from the Fix File is inserted into all selectd data files (Variable files).
We use this function to add detailed/renewed gene annotations to our ADF (Array Descriptions Files) or for building ADF files by merging generic spotter deconvolution files with our gene annotation database.

The example shows merging a genelist with a ChipSkipper data table using the "Reporter ID" column.
Select the Fix file (in our case the genelist).
The Fix file and variable file grids show partial previews ( about first hundred lines) of the selected files.
Click the respective column in the Fix File grid ("Link" shows up in column header) and the Variable files ("Link" shows up in column header) correspondingly.
the field in the link column of all selected
datafiles.
If no corresponding Link key is found in the "fix file"
nothing will happen to the data files.
Example:
Assume the Variable file:
ID_Number Comment1 ID_Char Comment2 Comment3 1 C1_1 A C2_1 C3_1 2 C1_2 B C2_2 C3_2 3 C1_3 C C2_3 C3_3 4 C1_4 D C2_4 C3_4 5 C1_5 E C2_5 C3_5 Assume the Fix-file
ID_Char New_Comment1 New_Comment2 B New_C1_B New_C2_B D New_C1_D New_C2_D E New_C1_E New_C2_E Use Link column ID_Char both in Variable as well as in Fix.
Replace Fix in Variable:
ID_Number Comment1 ID_Char New_Comment1 New_Comment2 Comment2 Comment3 1 C1_1 A C2_1 C3_1 2 C1_2 B New_C1_B New_C2_B C2_2 C3_2 3 C1_3 C C2_3 C3_3 4 C1_4 D New_C1_D New_C2_D C2_4 C3_4 5 C1_5 E New_C1_E New_C2_E C2_5 C3_5 Add Fix at the end of Variable:
ID_Number Comment1 ID_Char Comment2 Comment3 ID_Char New_Comment1 New_Comment2 1 C1_1 A C2_1 C3_1 2 C1_2 B C2_2 C3_2 B New_C1_B New_C2_B 3 C1_3 C C2_3 C3_3 4 C1_4 D C2_4 C3_4 D New_C1_D New_C2_D 5 C1_5 E C2_5 C3_5 E New_C1_E New_C2_E Add Fix in front of Variable:
ID_Char New_Comment1 New_Comment2 ID_Number Comment1 ID_Char Comment2 Comment3 1 C1_1 A C2_1 C3_1 B New_C1_B New_C2_B 2 C1_2 B C2_2 C3_2 3 C1_3 C C2_3 C3_3 D New_C1_D New_C2_D 4 C1_4 D C2_4 C3_4 E New_C1_E New_C2_E 5 C1_5 E C2_5 C3_5
Click the Rebuild files button to start the action.
Options:
Replace files = Source files are overwritten with merged data.
Empty Matchin lines = Ensures that renewed annotation is only appended to the
first occurence in the source list.
Reverse Mapping = Reverses the merging: Target files contain Source file
inserted into fix file.
All in one = A single file is created, where all source files are merged into one with the fix file as reference.
Assume you want to combine data files from different slide
batches of the same cloneset, but positions of single clones (i.e. lines in the
data files) have changed.
A simple way to combine such files would be to merge the data using the gene
name.
But a problem may arise if there are multiple copies of the same gene (e.g. GAPDH) on the array, resulting in lines with identical gene name in the data files.
TableButler inserts the FIRST occurence of a certain key (=gene name) to be placed into the variable data file. Thus, always the same version (e.g. first found version of GAPDH data line) will be inserted.
This ONLY way to solve this problem is to use a link column which contains UNIQUE feature-ids to be used as linking keys.
When starting table merging a dialog pops up:
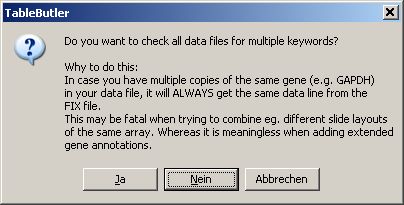
Select
If multiplicated lines are found, a messagebox pops-up:
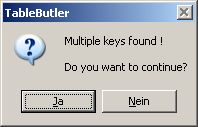
Click the Yes button to continue anyway. Click the NO button to cancel TableMerging.
A list of these keys is shown in the Log-tabsheet:
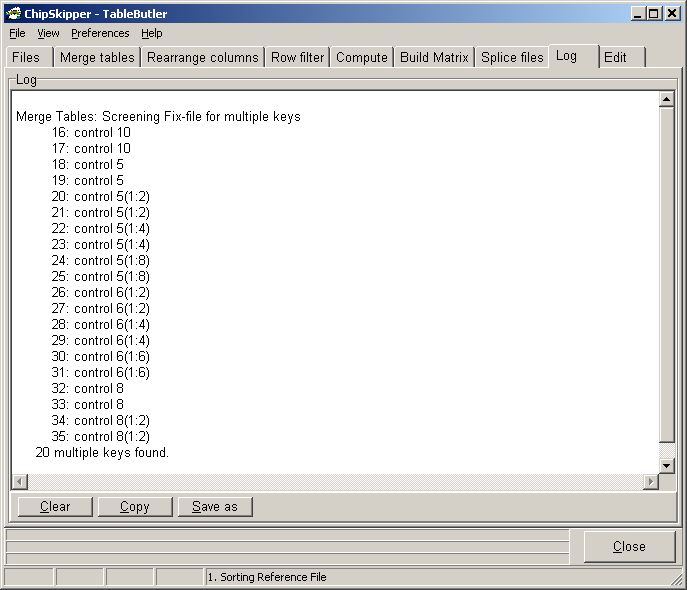
Last edited 10.06.2006,